Mapping
BFAST: A fast and accurate tool for mapping of short reads to reference sequences.
BWA: A fast light-weighted tool that aligns short nucleotide sequences to a sequence database.
Bowtie: An ultrafast, memory-efficient short read aligner.
ELAND: A very fast alignment algorithms from Illumina company.
MAQ: A software that builds mapping assemblies from short reads generated by the next-generation sequencing machines.
SHRiMP: A software package for aligning genomic reads against a target genome.
SOAP: A tool package that provides full solution to next generation sequencing data analysis (including a alignment tool SOAPaligner/soap2 etc).
SOLiD bioscope: A software package that is designed specifically to optimize the accuracy of the ABI SOLiD colorspace data.
SWIFT: A software collection for fast index-based sequence comparison.
TopHat: A spliced read mapper for RNA-Seq.
SNV Detection
CASAVA: The internal assembler and variant caller Illumina company utilized.
GATK: A multiple-sample, technology-aware SNV and indel caller.
JointSNVMix: A probabilistic model for detection of somatic mutations in normal/tumour pair.
SAMtools: A set of utilities for post-processing alignments in the SAM format, such as indexing, variant caller and alignment viewer.
SNVMix: A tool for SNV calling based on probabilistic binomial mixture model.
SOAPsnp: A tool for identifying SNVs by Beijing Genomics Institute (BGI).
Strelka: A tool for somatic small-variant calling from sequenced tumor-normal sample pairs.
SomaticSniper: A program to identify SNVs that are different between tumor and normal sample.
VarScan: A platform-independent, technology-independent software tool for identifying SNVs, indels, and CNVs in massively parallel sequencing of individual and pooled samples.
InDel Detection
Dindel: A program for calling small indels from short-read sequence data from Illumina platform.
Pindel: A tool for identifying indels and structural variants at single-based resolution from next-generation sequence data.
SplazerS: A tool for detecting genomic indel variants with exact breakpoints in single- and paired-end sequencing.
Structure Variation Detection
BreakDancer: A tool for detecting five types of SVs (insertions, deletions, inversions, inter- and intra-chromosomal translocations) from next generation paired-end sequencing reads.
CREST: A software that uses the soft-clipped reads to directly map the breakpoints of SVs.
GASV: A tool for identifying and comparing structural variants by computing intersections of breakpoint regions.
HYDRA: A tool for detecting structural variants in both unique and duplicated genomic regions.
PEMer: A software package for detecting SVs from paired-end reads.
R453Plus1Toolbox: An R/Bioconductor package for the analysis of Roche 454 sequencing data.
SVMerge: A tool for SVs analysis by integrating calls from several existing SV callers.
SVDetect: A tool for identifying structural variations from paired-end/mate pair data.
VariationHunter: An tool for identifying structural variations from paired-end WGS data.
Copy Number Variation Detection
CBS: An R package for detecting CNVs using sequencing data.
CMDS: A population-based method for recurrent CNVs analysis from multiple samples.
CNAseg: A tool for Identifying CNVs in cancer from NGS data.
cnvHMM: A tool for CNVs analysis using Hidden Markov algorithm.
CNVnator: A tool for CNV discovery and genotyping from depth of read mapping.
FREEC: A tool for control-free CNVs detection using deep-sequencing data.
RDXplorer: A tool for CNVs detection in whole human genome sequence data using read depth coverage.
SegSeq: A tool for detecting CNVs from short sequence reads.
VarScan: A platform-independent, technology-independent software tool for identifying SNVs, indels, and CNVs in massively parallel sequencing of individual and pooled samples.
Annotation
ANNOVAR: An efficient software tool to use update-to-date information to functionally annotate genetic variants detected from diverse genomes.
BreakSeq: A pipeline for annotation, classification and analysis of SVs at single nucleotide resolution.
Seattle Seq: An server that provides annotation of SNVs.
Data Visualization
Avadis: A software for visualizing and analyzing RNA-Seq data.
CIRCOS: A software package for visualizing genomic events.
IGV: A high-performance visualization tool for interactive exploration of next-generation sequencing data.
Pairoscope: A software package for generating diagrams indicating the relationship of paired end sequencing reads, is most useful for visualizing translocations.
UCSC Genome Browser: A genome browser that provide precise access to sequence and annotation data for any genomic region of specific interest.
Fusion Gene Detection
BreakFusion: A tool to identify gene fusions from paired-end RNA-Seq data.
Chimerascan: A software for detecting gene fusions in paired-end RNA sequencing (RNA-Seq) datasets.
Comrad: A tool to identify aberrant transcripts and associated rearrangements using low coverage genome data.
[defuse](http://sourceforge.net/apps/me ... _Page "Main_Page"): A software for discovering gene fusion from RNA-Seq data.
FusionAnalyser: A software for detecting gene fusions from paired-end RNA-Seq data.
FusionHunter: A tool to identify fusion transcripts from transcriptional analysis of paired-end RNA-seq.
FusionMap: A software to detect fusion junctions in both single- and paired-end dataset from either gDNA-Seq or RNA-Seq studies.
FusionSeq: A tool to identify fusion transcripts from paired-end RNA-sequencing.
nFuse: A tool for detecting fusion transcripts and associated complex genomic rearrangements from matched RNA-seq and whole genome shotgun sequencing.
SnowShoes-FTD: A tool to find fusions from RNA-Seq data.
ShortFuse: A software to identify fusions with ambiguously mapping read pairs without generating numerous spurious fusions from the many mapping locations.
TopHat-Fusion: A software with the ability to align reads across fusion points.
Trans-ABySS: A software to analyses ABySS-assembled contigs from shotgun transcriptome data.
来源:知因
为你推荐
 资讯
资讯 独家生产品种纳入国家基本药物目录应当经过单独论证,新版《国家基本药物目录管理办法》发布
国家基本药物工作协调机制由国家卫生健康委、国家发展改革委、工业和信息化部、财政部、商务部、市场监管总局、国家医保局、国家中医药局、国家疾控局、国家药监局和中央军委后...
2026-02-13 16:05
 资讯
资讯 玄言生物完成数千万元Pre-A轮融资,加速AI驱动精准诊疗产品落地
本轮融资资金将主要用于推进公司AI驱动的精准诊疗产品在多中心临床验证、注册申报及规模化应用中的落地进程,进一步夯实核心技术平台优势,拓展产品管线布局,助力AI医疗从科研...
2026-02-13 12:18
 资讯
资讯 赶早赴约,忆路守护 - 礼来携手清华大学启动阿尔茨海默病科普创意大赛
大赛旨在搭建汇聚青年智慧与专业力量的科普创作平台,鼓励以更具创意、更易传播的方式展开科学表达,推动公众减少对该疾病的误解,提升大众对于疾病和新出现的诊疗技术手段的理...
2026-02-13 12:08
 资讯
资讯 替尔泊肽单药治疗适应症获批,支持中国2型糖尿病长期管理获益
2026年2月11日,礼来中国宣布,穆峰达®(替尔泊肽注射液)获得中国国家药品监督管理局批准,用于成人2型糖尿病(T2DM)单药治疗。
2026-02-12 10:32
 资讯
资讯 智冉医疗完成3亿元A+轮融资,中科创星领投、老股东集体加注
本轮融资由硬科技投资领域头部机构中科创星领投,君联资本、北京市医药健康产业投资基金、IDG资本、顺为资本等一众老股东集体跟投加注
2026-02-12 10:28
 资讯
资讯 礼来安妥来利®安妥来®在华获批成人中重度活动性克罗恩病及溃疡性结肠炎双适应症
在针对克罗恩病(CD)的全球关键3期VIVID-1研究中,接受安妥来利®安妥来®治疗的患者在一年时临床缓解与内镜应答结局相较安慰剂呈显著改善
2026-02-11 15:37
 资讯
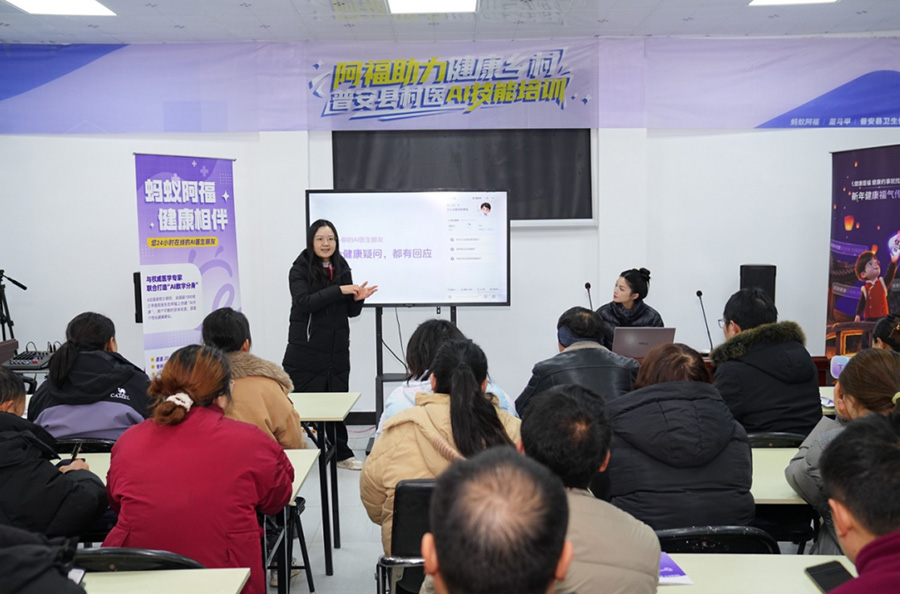
资讯 贵州普安将蚂蚁阿福引入村医一线工作
为了提升基层的平均诊疗水平,近期,普安县政府、卫健局等部门联合蚂蚁集团、阿里巴巴公益共同组织“村医AI技能培训”,教村医们学会用AI健康助手“蚂蚁阿福”来辅助日常的诊疗...
2026-02-10 18:01
 资讯
资讯 诺和诺德司美格鲁肽降糖注射版中国营收首次下滑
近日,诺和诺德正式发布2025年全年财报。报告显示,该公司2025年全年营收达3090 64亿丹麦克朗(约合489亿美元),同比增长6%;净利润为1024 34亿丹麦克朗(约合162亿美元),同比增长1%。
2026-02-09 14:23
 资讯
资讯 恒瑞医药HRS-4642注射液被纳入突破性治疗品种名单
2月6日,恒瑞医药(600276 SH)发布公告称,公司自主研发的HRS-4642注射液被国家药品监督管理局药品审评中心纳入突破性治疗品种名单。
2026-02-09 11:35
 资讯
资讯 强生达雷妥尤单抗注射液获批新适应症,四联疗法用于适合自体干细胞移植的新诊断多发性骨髓瘤成年患者的治疗
近日,强生宣布旗下兆珂速(达雷妥尤单抗注射液)获国家药监局批准拓展适应症,与硼替佐米、来那度胺和地塞米松联合,用于适合自体干细胞移植的新诊断多发性骨髓瘤成年患者的治疗。
2026-02-09 11:05
 资讯
资讯 信达生物与礼来达成合作,里程碑付款最高约85亿美元
根据合作协议,双方将发挥互补优势,加快推进创新药物的全球研发工作。信达生物依托自身成熟的抗体技术平台及高效的临床能力,将主导相关项目从药物发现至中国临床概念验证(二...
2026-02-08 20:55
 资讯
资讯 中国制药装备行业协会副秘书长遆倩鹤涉嫌严重违法,正接受监察调查
2月6日,中央纪委国家监委网站讯 据中央纪委国家监委驻中央社会工作部纪检监察组、海南省纪委监委消息:中国制药装备行业协会副秘书长遆倩鹤涉嫌严重违法,目前正接受中央纪委...
2026-02-07 22:13
 资讯
资讯 CDE:化学仿制药透皮和局部给药系统黏附性和刺激性/致敏性评估临床试验技术指导原则(试行)
本指导原则适用于TDS化学仿制药。常见适用TDS包括可能描述或称为贴片、透皮贴剂或缓释膜固态制剂产品。应用本指导原则时,请同时参考药物临床试验质量管理规范(GCP)国际人用药...
2026-02-07 13:48
 资讯
资讯 诺华可善挺(司库奇尤单抗)放射学阴性中轴型脊柱关节炎(nr-axSpA)新适应症在华获批
适用于治疗对非甾体类抗炎药(NSAID)应答不佳的活动性放射学阴性中轴型脊柱关节炎成人患者(其客观征象表现为C反应蛋白(CRP)升高和 或磁共振成像(MRI)证据)
2026-02-06 15:54
 资讯
资讯 天津市互联网诊疗监管实施办法(试行)
医疗机构应当主动与市级监管平台对接,及时上传、更新《医疗机构执业许可证》等相关执业信息,主动接受监督。医疗机构取得《医疗机构执业许可证》后或《医疗机构执业许可证》变...
2026-02-06 08:59